Lentiviral vectors (LVVs) have become the primary method of gene delivery for cells ex vivo in both academic and clinical applications. Currently, quantitative PCR (qPCR) is the gold standard for functional titering of LVVs, but Droplet Digital PCR (ddPCR) offers an attractive alternative due to its inherent benefits, such as absolute quantification without the use of a standard curve and enhanced resistance to PCR inhibitors. Ultimately, more precise titering will support the development of next-generation LVV-based therapeutics.
History of Lentiviral Vectors
The use of viral vectors to efficiently deliver genetic material to cells is an indispensable tool for both academic research and therapeutics. The idea of using a virus to deliver therapeutic genetic payloads was first realized with the development of γ-retroviral vectors in the 1980s and 1990s (Gutierrez-Guerrero et al. 2020). The first approved clinical trial was aimed at correcting severe combined immunodeficiency (SCID) through a γ-retrovirus–mediated gene transfer to hematopoietic stem cells in 1991 (Escors and Breckpot 2010). Although the therapeutic efficacy of the treatment is still heavily debated, the trial was deemed as a success in that the therapeutic was found to be safe.
In the early 2000s, two independent clinical trials demonstrated the promise of γ-retroviral therapy by introducing the common interleukin receptor γ-chain in bone marrow, and as a result corrected X-linked SCID in 11 children (Escors and Breckpot 2010). However, several trial participants went on to develop leukemia that was directly related to gene-insertion mutagenesis of the vector. Although this was deemed as a significant setback, development in other clinical applications continued to progress with γ-retroviruses.
In 2006, the first demonstration that immune cells could be genetically modified for cancer therapy was demonstrated in melanoma (Escors and Breckpot 2010). In that study, γ-retrovirus encoding the α- and β-chains of the melanoma antigen (MART-1)–specific T-cell receptor (TCR) was used to transduce peripheral blood lymphocytes, which were then administered back to patients. These autologous transduced T cells induced complete disease regression in 2 out of 15 patients.
Through the early development work and progression to clinical evaluation of γ-retroviruses, several observations, including the unexpected development of insertion mutagenesis–driven leukemia, highlighted the urgent need to better understand viral vector biology. It became clear that γ-retroviruses had specific limitations, including virus particle instability, inability to transduce resting cells, and a high rate of insertional mutagenesis, that would ultimately limit their clinical utility (Escors and Breckpot 2010). As a result, vectors derived from lentiviruses were developed to address some of these limitations. Unlike their γ-retrovirus predecessors, LVVs can efficiently transduce resting or nondividing cells and demonstrate a higher level of safety with a reduced risk of insertional mutagenesis due to their unique pattern of integration (Escors and Breckpot 2010, Milone and O’Doherty 2018). As a result of these advantages over the simpler γ-retroviruses, the use of LVVs has surged in recent years.
Lentivirus Structure and Function
Lentiviruses, such as human immunodeficiency virus (HIV), belong to the family Retroviridae (Segura et al. 2006). LVVs are 80–120 nm in diameter and contain two copies of a positive-sense RNA genome (Gouvarchin et al. 2020). The RNA genome is complexed with nucleocapsid protein, along with a reverse transcriptase, integrase, and protease (Escors and Breckpot 2010). Matrix proteins form the external layer and interact with the host cell–derived lipid envelope. This lipid envelope contains glycoproteins derived from the env gene that interact with specific host cell receptors (Escors and Breckpot 2010). The genome of lentiviruses, like all retroviruses, contains the gag, pol, and env genes, which are flanked by long terminal repeats (LTRs) (Gouvarchin Ghaleh et al. 2020). In addition, lentiviruses also contain vif, vpr, vpu, nef, tat, and rev, which serve as both accessory and regulatory genes and encode proteins that have roles in infection, binding, and release (Figure 1) (Gouvarchin Ghaleh et al. 2020).
Similar to the HIV life cycle, LVV-based gene delivery requires cell surface binding and subsequent internalization to the host cell. However, wild type HIV has a severely limited subset of cells that it can bind to and subsequently infect. To enhance the variety of cell types that would be amenable to LVV-mediated transduction, LVVs are pseudotyped by exchanging the env gene of the virus, effectively swapping out the envelope. The most popular pseudotype uses the env gene from vesicular stomatitis virus (VSV-G), though other viral envelopes have also been utilized (Coil and Miller 2004). LVVs pseudotyped with VSV.G exhibit significant advantages, such as enhanced vector stability, relative to wild type (Breckpot et al. 2007). Furthermore, VSV.G interacts with a ubiquitously expressed cell surface receptor that greatly enhances cells’ receptivity to LVV transduction (Coil and Miller 2004).
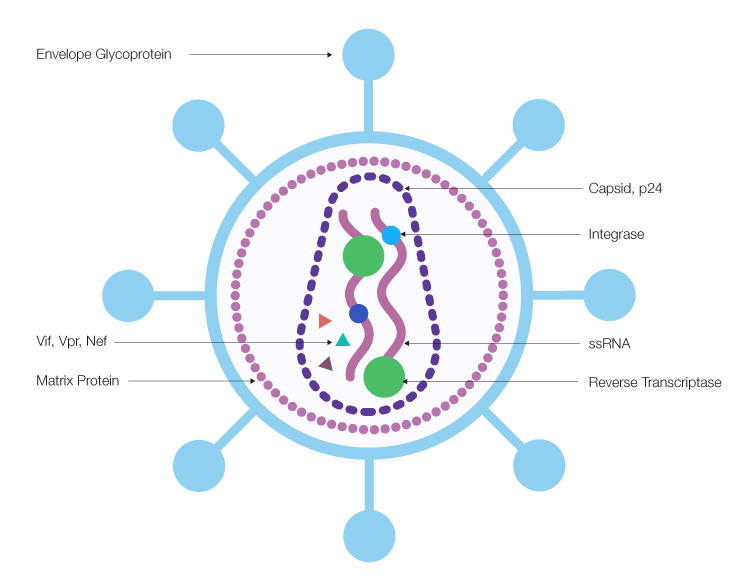
Fig. 1. Lentivirus structure and composition.The positive-sense single-stranded RNA (ssRNA) genome is packaged in a capsid with viral proteins, including Nef, Vpr, and Vif accessory proteins, reverse transcriptase, and integrase. The capsid is surrounded by a shell comprised of matrix protein, then an outer membrane dotted with envelope glycoproteins.
Clinical and Research Applications of Lentiviral Vectors
Due to their high transduction efficiency and integrative capacity, LVVs are the preferred vector for ex vivo gene correction (Escors and Breckpot, 2010). Currently there are more than 100 ongoing clinical trials in the United States (U.S.), China, European Union (EU), and Canada that incorporate LVVs for the correction of monogenic diseases affecting the hematopoietic system. Recently, the LVV-based gene therapy Zenteglo received regulatory approval in the U.S. for the treatment of transfusion-dependent β-thalassemia (Asghar et al. 2022). This autologous LVV-transduced hematopoietic stem cell product is potentially curative through the introduction of a b-globin gene to improve the production of normal hemoglobin. Further, extensive preclinical and clinical evaluation of similar approaches are underway for X-linked adrenoleukodystrophy, metachromatic leukodystrophy, and Wiskott-Aldrich Syndrome (Milone and O’Doherty 2018, Cavazzana-Calvo et al. 2010, Eichler et al 2017).
LVVs are also used extensively in transducing T cells with novel chimeric antigen and engineered T-cell receptors for cancer therapy (Labbé et al. 2021). The commercially approved T-cell products Yescarta, Kymriah, and Tecartus all utilize receptors delivered by LVVs (Labbé et al. 2021, Perry and Rayat 2021). Furthermore, recent years have witnessed an exponential growth in the number of clinical trials utilizing autologous T-cell products transduced by LVVs to express novel receptor constructs and numerous tumor antigens. In addition, LVVs have been utilized to transduce dendritic cells ex vivo to elicit strong T-cell responses to specific tumor antigens in vivo (Breckpot et al. 2007, Kim et al. 2017).
Finally, LVVs have been broadly used in research applications for gene editing and modulation of gene expression. Regarding their use in gene editing, LVV have been shown to be extremely efficient delivery vehicles for the CRISPR-Cas9 gene editing system, which is capable of excising, adding, or editing specific genes (Rittiner et al. 2022). In the context of gene modulation, cells in culture have been transduced with LVVs carrying a variety of inducible and adjustable expression cassettes, such as one-piece inducible autoregulatory cassettes (Ogueta et al. 2001). Similarly, a variety of cell lines have been transduced with LVVs carrying streptogramin–adjustable expression of reporter differentiation–modulating transgenes (Mitta et al. 2004).
Challenges Unique to Lentiviral Vectors
LVVs are produced in cell culture via transient transfection of the required plasmid sets into a packaging cell line, such as HEK 293T (Milone and O’Doherty et al. 2018). Plasmid sets for LVVs have been categorized into generations according to the configuration of the packaging plasmid (Breckpot et al. 2007). However, in general, cotransfection of plasmids required for LVV production includes a packaging plasmid, a transfer plasmid that contains the gene of interest, and an envelope plasmid (Breckpot et al. 2007).
During the cell culture phase of production, LVVs are secreted into the cell culture medium, where they can be harvested, although subsequent purification is required for use in clinical applications. For clinical purposes, it is necessary to purify LVVs to remove product- and process-derived impurities, such as host cell DNA and proteins, residual plasmid DNA, degraded or nonfunctional vectors, and cell culture media contaminates, to achieve acceptable purity (Labbé et al. 2021). Purification of LVVs can be inherently complex and challenging due to sensitivity of the vector to several physical or environmental conditions typically encountered during the purification and subsequent storage process (Higashikawa and Chang 2001, Kumru et al 2018). Care must be taken to determine viral quantity and quality through downstream unit operations to ensure minimization of vector loss and to maintain high functional quality.
In addition to purification, the LVVs produced must be quantified. Accurate quantitation of functional virus is critical for dosing, as an excessively high multiplicity of infection (MOI) can have adverse effects on the host cell to be transduced. LVVs can be quantified using either physical or functional titering methods.
Physical titering, which measures the number of virus and virus-like particles, is typically done by enzyme-linked immunosorbent assay (ELISA)-based detection of p24, the most abundant HIV protein (Perry and Rayat 2021). Although physical titering is straightforward, it cannot discriminate between functional and nonfunctional particles, unprocessed p24, and degraded vector, which can lead to drastic overestimation of viral titer. Furthermore, these physical titering kits rely on the specificity of the antibodies utilized, so titers may not be reproducible across different vendors (Perry and Rayat 2021).
Functional titering, which measures the quantity of virus particles able to transduce target cells, can be achieved by several different methodologies. Flow cytometry (FCM) of transduced cells is achievable if the LVV-transduced gene contains a fluorescent reporter or if the transgene product can be labeled using staining antibodies. Antibiotic resistance markers have also been known to be used. Measures of functional titer produce a percentage of a target cells that have been determined to be transduced. However, FCM cannot discriminate between single or multiple infection events per cell, which can greatly affect the titering calculation (Wang et al. 2018).
Determination of lentivirus proviral copy number in the host genome of transduced cells by qPCR is the standard method used for LVV functional titer quantification (Wang et al 2018, Gill and Denham 2020). However, accurate titering is dependent on the availability of a reference LVV with which to generate a standard curve (Gill and Denham 2020). The standard curve is generated by performing multiple infections with serially diluted reference standard in biological triplicate and subsequently determining the linear portion of the standard curve. This is both time consuming and potentially imprecise, given the inherent variability of cell culture. Furthermore, the results of qPCR are highly dependent on the quality and composition of the sample, as qPCR is susceptible to inhibition by commonly present chemicals and contaminates (Wang et al. 2018).
The inherent benefits of ddPCR technology make it an attractive alternative to qPCR for LVV functional titering and other cell and gene therapy applications. Specifically, in ddPCR assays, each sample is partitioned into tens of thousands of tiny droplets with a known volume and each droplet is subjected to PCR amplification. Droplets are classified as either positive or negative for the target, then the target DNA copy number in the original sample is determined using Poisson statistics (Wang et al. 2018). This method offers several advantages over qPCR, such as enabling LVV titering without the use of a standard curve. ddPCR technology is also less sensitive to PCR inhibitors, relative to qPCR, that may be present in the sample matrix. As a result, LVV titering that utilizes ddPCR assays can offer simplified and straightforward workflows that are more sensitive and precise.
Future Direction and Closing Remarks
LVVs offer several benefits over γ-retrovirus vectors and have become the primary method of gene delivery for cells ex vivo in both academic and clinical applications. As the evolution of gene therapy continues to progress, the use and scope of these vectors will only increase. LVVs are extremely efficient at gene delivery. However, LVVs demonstrate inherent particle sensitivity and have been known to lose functionality when subjected to environmental and physical stresses. This instability can generate interfering components, which can lead to extremely misleading titering results with specific methods. Thus, determining functional quality is of the utmost importance for both downstream process development as well as determination of dose for research and clinical applications.
Currently, qPCR is considered the gold standard for functional titering of LVVs, but qPCR workflows are not ideal in that they require a standard curve using a reference standard and they suffer from the method’s susceptibility to PCR inhibitors. Technologies utilized in the development, manufacture, and release of these vectors must progress within the LVV space to adequately support the progression of the technology. The application of ddPCR technology offers an excellent alternative to qPCR and other assays due to its inherent benefits, which include absolute quantification without the use of a standard curve and enhanced resistance to PCR inhibitors. Functional titering using ddPCR technology can provide more accurate and sensitive LVV quantification than other methods. Ultimately, more precise titering will support the development of next-generation LVV-based therapeutics to offer better treatment options for patients with previously untreatable conditions.
Bio-Rad offers a comprehensive portfolio of ddPCR instruments, consumables, reagents, and kits to support virtually any application.
References
Asghar AA et al. (2022). Zynteglo: Betibeglogene autotemcel – An innovative therapy for β- thalassemia patients. Ann Med Surg (Lond) 13, 104624.
Breckpot K et al. (2007). Lentiviral vectors for cancer immunotherapy: Transforming infectious particles into therapeutics. Gene Ther 14, 847–862.
Cavazzana-Calvo M et al. (2010). Transfusion independence and HMGA2 activation after gene therapy of human β-thalassaemia. Nature 467, 318–322.
Coil DA and Miller AD (2004). Phosphatidylserine is not the cell surface receptor for vesicular stomatitis virus. J Virol 78, 10920–10926.2004.
Eichler F et al. (2017). Hematopoietic stem-cell gene therapy for cerebral adrenoleukodystrophy. N Engl J Med 377, 1630–1638.
Escors D and Breckpot K (2010). Lentiviral vectors in gene therapy: Their current status and future potential. Arch Immunol Therp Exp (Warsz) 58, 107–19.
Gill KP and Denham M (2020). Optimized transgene delivery using third-generation lentiviruses. Curr Protoc Mol Biol 133, e125.
Gouvarchin Ghaleh HE et al. (2020). Concise review on optimized methods in production and transduction of lentiviral vectors in order to facilitate immunotherapy and gene therapy. Biomed Pharmacother 128, 110276.
Gutierrez‐Guerrero A et al. (2020). Lentiviral vector pseudotypes: Precious tools to improve gene modification of hematopoietic cells for research and gene therapy. Viruses 12, 1016.
Higashikawa F and Chang L (2001). Kinetic analyses of stability of simple and complex retroviral vectors. Virology 280, 124–131.
Kim JT et al. (2017). Dendritic cell–targeted lentiviral vector immunization uses pseudotransduction and DNA-mediated STING and cGAS activation. Sci Immunol 2, eaal1329.
Kumru OS et al. (2018). Physical characterization and stabilization of a lentiviral vector against adsorption and freeze-thaw. J Pharm Sci 107, 2764–2774.
Labbé RP et al. (2021). Lentiviral vectors for T cell engineering: Clinical applications, bioprocessing and future perspectives. Viruses 13, 1528.
Milone MC and O’Doherty U (2018). Clinical use of lentiviral vectors. Leukemia 32, 1579–1541.
Mitta B et al. (2004). Design and in vivo characterization of self-inactivating human and non-human lentiviral expression vectors engineered for streptogramin-adjustable transgene expression. Nucleic Acids Res 32, 1–14.
Ogueta SB et al. (2001). Design and in vitro characterization of a single regulatory module for efficient control of gene expression in both plasmid DNA and a self-inactivating lentiviral vector. Mol Med 7, 569–579.
Perry C and Rayat ACME (2021). Lentiviral vector bioprocessing. Viruses 13, 268.
Rittiner J et al. (2022). Therapeutic modulation of gene expression in the disease state: Treatment strategies and approaches for the development of next-generation of the epigenetic drugs. Front Bioeng Biotechnol 10, 1035543.
Segura MdlM et al. (2006). Purification and characterization of retrovirus vector particles by rate zonal ultracentrifugation. J Virol Methods 133, 82–91.
Wang Y et al. (2018). Determination of lentiviral infectious titer by a novel Droplet Digital PCR method. Hum Gene Ther Methods 29, 96–103.