In nature, mammalian extracellular vesicles (EVs) are vital for intercellular communication, immune responses, and cell removal. EVs also hold promise in diagnostics, therapeutics, and vaccine delivery due to their abundance and natural role as carriers. Explore how overcoming challenges in EV purification and precise RNA analysis methods can help unlock their potential applications.
- Introduction
- Multiple Isolation Strategies Are Required for EVs
- Molecular Analysis of EVs for Gene Expression Profiling
- Emerging Technologies and Future Directions
- Conclusion
- References
Introduction
The Importance of Extracellular Vesicles (EVs) in Intercellular Communication
Secreted by virtually all mammalian cell types, EVs play crucial roles in intercellular communication by transporting bioactive molecules such as proteins, nucleic acids, and lipids. The spectrum of interest in EVs ranges from understanding their basic biology to their application in diagnostics as biomarkers, in addition to cell-free delivery systems for therapeutics and vaccines (Ilahibaks et al. 2024).
EVs and liquid nanoparticles (LNPs) are both valuable carriers for therapeutics, with LNPs emerging as alternatives to EVs, especially in vaccine development, due to their efficient loading and consistent production. However, choosing between them depends on the therapeutic goals, with challenges existing in both methods (Maugeri 2019). EV research has spawned diverse applications beyond vaccines, including regenerative medicine, cancer therapeutics, and drug delivery. EVs possess natural targeting abilities, low immunogenicity, and the capacity to transport various cargo molecules, potentially serving where mRNA vaccines are suboptimal. Despite the success of mRNA vaccines, EV research remains crucial due to its promising applications and therapeutic benefits, necessitating ongoing exploration and investment.
Challenges in EV Isolation and Molecular Analysis of RNA Transcripts
EVs represent a complex and heterogenous group of vesicles, including microvesicles, exosomes, and apoptotic bodies. In addition to variations in size, the site of origin of EVs can also alter the membrane composition and cargo. Due to this complexity, as well as the overlapping size distribution with other extracellular particles and the presence of contaminants with similar size properties or non-EV lipid particles and intracellular proteins, EV purification is fraught with challenges (Newman et al. 2023).
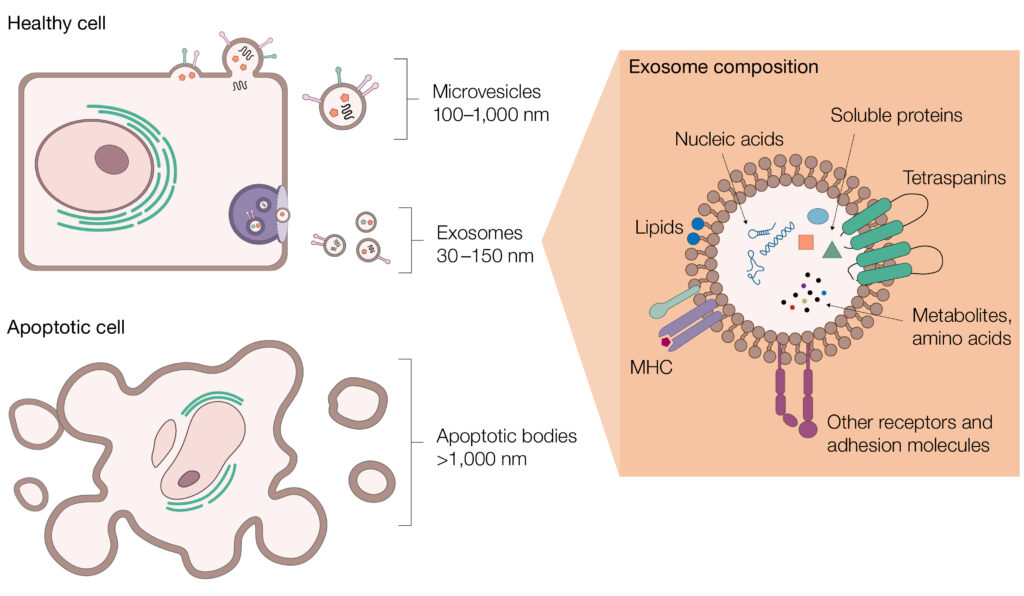
Extracellular vesicles (EVs) are heterogenous and complex. EVs include microvesicles, exosomes, and apoptotic bodies. Each of these vesicle types have different sizes, contents, and surface proteins, making their isolation challenging.
These purification challenges can make it difficult to obtain EVs of sufficient quantity and quality for further analysis and in a reproducible manner, which directly impacts the reliability of downstream conclusions (Nieuwland and Siljander 2024).
The challenges of low purity and reproducibility in current EV isolation methods may also affect the subsequent molecular analysis of EV RNA transcripts. Moreover, the analysis of RNA transcripts from EVs presents the risk of RNA degradation during isolation and the problem of low-abundance targets. To overcome these, precise sample handling and sensitive detection methods like Droplet Digital PCR (ddPCR) or RNA sequencing (RNA-Seq) are vital. To achieve reliable results, standardized EV isolation protocols, implementation of quality control measures, and refining analysis methods are essential and can aid in demonstrating functional RNA transfer in living cells.
Addressing these obstacles is vital to advance our understanding of EV biology and unlock their therapeutic potential. Such advancements will also be accompanied by progress in proteomic or lipidomic analysis.
Multiple Isolation Strategies Are Required for EVs
Researchers currently have at their disposal an array of isolation techniques that capitalize on various biophysical or biochemical characteristics of EVs (Zeringer et al. 2015). However, as mentioned above, isolating EVs from biological fluids or cell culture supernatants faces numerous challenges and no standalone one-size-fits-all technique exists for this purpose.
In addition to issues with complexity, heterogeneity, and overlapping properties with other particles, EV purification is further hindered by the range of possible contaminants, which can change depending on the source of EVs. For example, lipoproteins are highly abundant in blood and have a similar size and density as exosomes, making isolation techniques that solely rely on these properties ineffective (Simonsen 2017). In addition to lipoproteins, viruses also can have a similar size and density as EVs and may co-purify with them during isolation, necessitating additional quality control steps (Reiter et al. 2019). Equally, the presence of endotoxin can be a concern and may hinder subsequent experiments or even mask biological effects (Babula et al. 2023). Thus, the choice of which isolation technique to employ hinges on the study aims and the sample complexity — in short, the isolation strategy must match the application.
Techniques for EV Isolation
EV isolation techniques can be broadly divided into five categories — density, solubility, size, charge, and affinity-based techniques. Each has its specific advantages and disadvantages, and no single technique can achieve comparable purity and yield levels across the many different sample types. A balance must be achieved between yield and purity, and researchers should be aware of their chosen isolation technique’s limitations to adjust their conclusions accordingly. When reviewing literature, researchers should also be aware that the terms isolation and purification are frequently used in contexts where EVs are merely enriched or concentrated. Even though absolute purification of EVs is an unattainable goal, their purity can be increased by combining various purification techniques (Théry et al. 2018).
The current landscape of EV isolation/separation techniques has been discussed in numerous articles (Liangsupree et al. 2020, Clos-Sansalvador et al. 2022, Akbar et al. 2022). In Table 1, we summarize the principles, benefits and limitations of the main techniques and their primary use(s). In general, it is difficult to compare the performance of different techniques as comparative studies are rare. In addition, there are many sample types, volumes, and purity requirements that also influence the usability of a specific technique. Thus, a technique that may perform well for a given application may not yield adequate/satisfactory results for another application.
Table 1. Principles, benefits, and limitations of most common EV isolation techniques.
| Technique | Principle | Benefits | Limitations |
| Affinity Chromatography | Relies on the selective and specific interaction between EV surface exposed components and ligands immobilized on a support (such magnetic beads or chromatography resins). Most commonly, the immobilized ligands are capturing antibodies, but other ligands such as heparin or peptides exist. |
|
|
| Microfluidics | Emerging microscale technique which, depending on the specific microfluidics platform used, can isolate EVs using one or a combination of different separation mechanisms such as affinity, size or density, or novel sorting mechanisms such as acoustic, electrophoretic, or electromagnetic fields. |
|
|
| Technique | Principle | Benefits | Limitations |
| Precipitation | Based on altering EV solubility, often by use of hydrophilic polymers such as polyethylene glycol (PEG). By competing/tying up water molecules, the PEG forces less-soluble components such as EVs out of solution, which can be then collected by a low-speed centrifugation step. |
|
|
| Ultrafiltration (UF) | Separation based on size differences between EVs and other sample components. Relies on the use of membranes with specific pore sizes to isolate sample components of a predetermined size range. Typically, components larger than EVs are first filtered out, resulting in a retentate containing EVs. The retentate is subjected to a second filtration step using a membrane with pores smaller than EVs, such that smaller contaminants pass into a waste eluate. |
|
|
| Tangential flow filtration (TFF) | As in the case for UF, separation relies on size. A feed stream flows, in contrast to UF, parallel to a membrane with a pore size smaller than EVs. Non-EV components continuously pass through the membrane and end up in the waste, while EV containing retentate is circulated back to the sample reservoir numerous times, such that EV sample gets concentrated with each round. |
|
|
| Technique | Principle | Benefits | Limitations |
| Ion Exchange Chromatography (IEX) | Exploits the net negative charge found on the EV surface. Can take two forms, using either an anion exchange (AEX) resin or a cation exchange (CEX) resin. In the AEX method, EV particles bind to a positively charged resin (anion exchanger). Bound EVs are then desorbed or eluted by increasing the ionic strength of the buffer surrounding the resin. In contrast, negatively charged resins (cation exchangers) can be used to bind and remove contaminants to increase EV purity. |
|
|
| Technique | Principle | Benefits | Limitations |
| Size Exclusion Chromatography (SEC) | Separation based on the size differences between EV particles and other sample components. Uses a chromatography resin with pores of a specific size distribution. EVs are generally much larger than pores of the resin, thus are excluded from entry and quickly elute, whereas contaminants enter and travel through the resin resulting in much later elution. |
|
|
| Differential Ultracentrifugation (dUC) | Sequential separation of EV particles from other sample components based on their size and density using a series of centrifugal forces and durations. EVs are pelleted in the final centrifugation step. |
|
|
| Density Gradient Centrifugation (DGC) | Separation based on size and density similar to dUC; however, separation occurs over preconstructed density gradient (typically made of sucrose or iodoxinol). EVs are segregated to specific layers according to their buoyancy. |
|
|
Size Exclusion Chromatography (SEC)
SEC has gained recognition as a valuable method in EV isolation (Sidhom et al. 2020). This is because SEC offers a reproducible technique capable of isolating intact EVs at high yield and purity. EV isolations by SEC are generally conducted using a gravity flow column, requiring significant experimenter involvement and contradicting efforts towards the standardization and reproducibility desired by the EV research community.
SEC can be performed manually or using an NGC Chromatography System, which enables a controlled process and real-time monitoring of EV separations at different wavelengths, including those used to monitor labeled EVs. Small volumes collected into 96-well plates with the NGC Fraction Collector or the BioFrac Fraction Collector can facilitate the collection of purer EV-containing fractions ready for downstream analysis. Automation of EV separations, standard sample runs, column QC, and column cleaning and maintenance can all be achieved with the NGC Chromatography System.
Capture and Concentration of EVs
SEC following filtration-based methods is a common trend in numerous publications — particularly ultrafiltration, a form of dead-end filtration. Although some reduction in contaminants occurs, filtration mainly serves the purpose of concentrating the sample prior to SEC. Concerns with ultrafiltration are EV loss on the membrane and formation of protein aggregates, which may carry through to the subsequent size-based EV isolation. Tangential flow filtration (TFF) may not suffer from these drawbacks; however, it is not readily available to most researchers.
An alternative to filtration is the precipitation of EVs. Although simple to perform, precipitation is difficult to standardize in terms of reproducibility and data acquisition and is not considered a scalable solution, especially for therapeutic applications. In addition, precipitation could diminish the biological activity of EVs as previously reported (Paolini et al. 2016).
Combining SEC with chromatography techniques such as affinity chromatography could solve some of these problems. In principle, affinity chromatography is an appealing workflow step as it could provide an already highly pure and concentrated sample before polishing by SEC, however, current technology based on immunoaffinity is costly and not considered scalable (Table 1). Newer affinity ligands are currently being explored which may solve this limitation (Benayas et al. 2023). Heparin affinity chromatography has recently been proposed as an inherently scalable method, but its utility is not for capture but rather as part of a multistep workflow to improve purity (Zhou et al. 2023).
EV Capture using Anion Exchange Chromatography (AEX)
Another scalable chromatography technique is AEX, which is frequently used to increase EV purity after TFF. AEX exploits negative charges on the surface of EVs and their interactions with a positively charged chromatographic matrix. However, AEX can also be used to rapidly capture and concentrate EVs in place of ultrafiltration or TFF (Seo et al. 2022, Malvicini et al. 2023, Koch et al. 2024). AEX can provide decreased contamination as compared to TFF-concentrated EVs (Heath et al. 2018).
One challenge when working with large biomolecules, such as EVs, is access to the chromatography resin’s absorption sites, which are not only located on the surface but, more importantly, within the resin accessible through the pores. If EVs are not able to reach the absorption sites, because resin pores are not large enough, this would result in inefficient binding and an overall low capacity of the resin to bind EVs. An AEX resin such as Nuvia HP-Q Resin, specifically designed for large biomolecules, is an attractive candidate for efficient capture and release of EVs.
A capture step from conditioned media using Nuvia HP-Q Resin could be complemented by an SEC step using Bio-Gel A-1.5m Gel to establish a highly reproducible and automatable chromatography workflow using the NGC Chromatography System.
Molecular Analysis of EVs for Gene Expression Profiling
Techniques: RT-qPCR, ddPCR, RNA Sequencing
Once exosomes and/or EVs have been isolated and purified, it is useful to characterize them to gain insights into their functions. Various biomolecules, including RNA, are contained within EVs when they are released into the extracellular environment. Thus, gene expression analysis of EV RNA transcripts can provide valuable insights into their roles in cellular communication and disease processes.
Several techniques are commonly employed for the purpose of molecular analysis:
- Reverse transcription quantitative PCR (RT-qPCR) — widely used for the quantitative analysis of RNA transcripts in EVs, as it offers high sensitivity, specificity, and a wide dynamic range. It involves reverse transcription of RNA into complementary DNA (cDNA), followed by quantitative PCR (qPCR) to amplify and quantify specific target genes (Gouin et al. 2017)
- Droplet Digital PCR — partitions the PCR reaction mixture into thousands of individual droplets, allowing for absolute quantification of target RNA molecules. ddPCR technology offers higher precision and sensitivity compared to qPCR, making it particularly useful for detecting rare or low-abundance transcripts in EVs (Jia et al. 2014)
- RNA sequencing (RNA-Seq) — a high-throughput technique for profiling the EV transcriptome, especially the small RNAs. It involves sequencing cDNA synthesized from EV RNA transcripts, followed by bioinformatic analysis to identify and quantify differentially expressed genes. The comprehensive nature of RNA-seq gene expression profiling allows for the discovery of novel transcripts and splice variants in EVs (Conley et al. 2016)
- Long-read RNA-Seq — uses specialized system; can generate sequence reads that span entire RNA transcripts, including full-length isoforms and splice variants. These types of RNA processing events may be missed or inaccurately reconstructed by short-read sequencing methods. Thus, this capability of long-read sequencing can provide a more comprehensive and accurate representation of the EV transcriptome than short-read technology (Rodosthenous et al. 2020)
Challenges in RNA Transcript Analysis
EV gene expression analysis is faced with several challenges, including:
- Degradation of RNA — RNA molecules from EVs may degrade during isolation and analysis. Careful handling and storage of EV samples to preserve their integrity are essential to minimize RNA degradation and prevent loss of valuable information (Buschmann et al. 2017)
- Low-abundance targets — EVs contain a heterogeneous mixture of RNAs, including low-abundance transcripts that can be overlooked by traditional techniques. Sensitive detection methods, such as ddPCR or RNA-seq, are required to accurately quantify low-abundance targets in EVs (Jia et al. 2014)
- Other challenges — Research on EV RNA encounters numerous technical obstacles, including the potential for contamination from non-vesicular RNA sources and the need to standardize EV isolation techniques in terms of protocols and quality control measures to ensure reproducibility and reliability of results. Moreover, researchers have noted interest in how to refine methods for isolating and analyzing small amounts of RNA contained within EVs and devise strategies to showcase the functional transfer of EV RNA within living organisms (Mateescu et al. 2017, Yi et al. 2023)
Key Findings from Molecular Analysis of EVs and Their Implications/Significance:
EV gene expression analysis offers key opportunities for advancing biomedical research and clinical applications. Recent findings have unveiled several important insights into the role of EVs in intercellular communication, disease mechanisms, and their therapeutic potential. Despite the challenges associated with these techniques, the opportunities for biomarker discovery and disease diagnosis using EV gene expression analysis are substantial, paving the way for personalized medicine and improved patient care. A summary of these core insights provided by analysis of EVs is described in Table 2.
Table 2. Opportunities for the use of EV gene expression analysis.
| Application | Insights | Emerging Use Cases |
| Biomarker discovery |
EVs contain diverse RNA types, includes microRNAs, mRNAs, and long non-coding RNAs. The population of RNAs in EVs is indicative of the physiological or pathological state of their parental cells. Thus, profiling the EV gene expression signature enables the discovery of novel biomarkers for disease diagnosis, prognosis, and therapeutic response monitoring. |
|
| Disease detection and monitoring |
Alterations in EV gene expression profiles have been implicated in various diseases, including cancer, neurodegenerative disorders, and cardiovascular diseases. Notably, researchers have identified specific RNA transcripts carried by EVs that serve as biomarkers for these various diseases. Use of these biomarkers can potentially enable early disease detection, prognosis assessment, and treatment monitoring. |
Liquid biopsies are an emerging source of EV-based biomarkers in oncology, providing a new dimension in disease management by reflecting their cancer cell origins through selectively sorted cargos (Rayamajhi 2024). These EV-based biomarkers demonstrate notable performance across cancer stages, facilitating early detection, therapy response prediction, disease burden tracking, and cancer recurrence identification. Additionally, liquid biopsies offer distinct advantages over traditional biopsies, including non-invasiveness, precision, and high sensitivity, potentially surpassing traditional diagnostics (Lianidou and Pantel 2019). This underscores the promising potential of EVs from liquid biopsies to revolutionize cancer monitoring and enhance patient care, emphasizing the need to address contaminant challenges for meticulous research-driven clinical translation (Irmer et al. 2023). |
| Application | Insights |
| Cell-to-cell communication | EVs have been found to transfer functional mRNA and microRNA molecules between cells, regulating gene expression and influencing recipient cell behavior. This communication mechanism plays crucial roles in physiological processes such as tissue homeostasis, immune response modulation, and neuronal signaling (Sork 2018). The selective sorting of microRNAs into recipient EVs may influence many diseases, including diabetes, cancer, and heart disease (Groot 2020). |
| Therapeutic cargo delivery | EVs loaded with therapeutic nucleic acids, such as small interfering RNAs (siRNAs) or microRNAs, have shown promise as delivery vehicles for gene therapy. Small EV mimetics have been proposed as well for therapeutic delivery of RNA molecules, for example (Geng 2021). Gene expression analysis helps identify the most effective cargo for targeting specific cellular pathways or correcting genetic defects. |
| Tumor heterogeneity and drug resistance | EV-mediated transfer of RNA molecules has been implicated in promoting tumor heterogeneity and drug resistance by facilitating communication between different tumor cell populations or between tumor cells and the tumor microenvironment. Understanding the gene expression profiles of EVs sheds light on these mechanisms and offers opportunities for developing targeted therapies. |
| Crosstalk between tumor cells and stroma | Gene expression analysis of EVs derived from tumor cells has revealed their role in modulating the tumor microenvironment, including interactions with stromal cells, immune cells, and endothelial cells. This crosstalk influences tumor progression, angiogenesis, and metastasis. |
Emerging Technologies and Future Directions
The field of EV research faces challenges due to a lack of standardization in data generation, acquisition, and reporting. Efforts by the International Society for Extracellular Vesicles task force, such as the MISEV 2023 guidelines, aim to accelerate scientific progress in this area. Tools like EV-TRACK and recombinant extracellular vesicles serve to further enhance research speed (Geeurickx et al. 2019).
However, to fully utilize EVs for diagnostics and therapeutics, their separation must be addressed in terms of automation, throughput, and reproducibility. Chromatography presents a solution to these challenges, facilitating standardization and scalability for EV therapies. Technological advancements are crucial for enabling robust, high-throughput isolation workflows without compromising purity or yield, especially when combined with multi-omics analysis to understand biomarkers better. While microfluidic devices show promise in addressing throughput issues, their current limitations in sample recovery raise concerns regarding robust analysis using various -omics approaches.
Conclusion
In summary, the challenges and opportunities surrounding EVs in biomedical research underscore the necessity of overcoming technical hurdles to harness their full potential. Despite the complexities of EV isolation and molecular analysis, advancements in technology and opportunities for interdisciplinary collaboration offer promising avenues to address these challenges. Pooling expertise from diverse fields such as biology, engineering, and medicine can aid in the development of innovative techniques for EV isolation and molecular analysis, and ultimately provide standardized protocols for reproducible results.
The potential of EVs to revolutionize biomedical research and clinical applications is vast, with the prospect of transforming disease diagnosis, prognosis, and therapy. Unraveling the complexities of EV biology and their functions in intercellular communication presents a promising future landscape in which EVs act as potent vehicles for precision medicine and tailored therapeutics, improving comprehension of disease processes and ultimately enhancing patient outcomes.
References
Akbar A et al. (2022). Methologies to isolate and purify clinical grade extracellular vesicles for medical applications. Cells 11, 186.
Babula A et al. (2023). Isolation of low endotoxin content extracellular vesicles derived from cancer cell lines. J Vis Exp 192, e65062.
Benayas B et al (2023). Proof of concept of using a membrane-sensing peptide for sEVs affinity-based isolation. Front Bioeng Biotechnol 11, 1238898.
Buschmann D et al. (2018). Evaluation of serum extracellular vesicle isolation methods for profiling miRNAs by next-generation sequencing. J Extracell Vesicles 7, 1481321.
Clos-Sansalvador M et al. (2022). Commonly used methods for extracellular vesicles’ enrichment: Implications in downstream analyses and use. Eur J Cell Biol 101, 151227.
Conley A et al. (2017). High-throughput sequencing of two populations of extracellular vesicles provides an mRNA signature that can be detected in the circulation of breast cancer patients. RNA Biol 14, 305–316.
Geeurickx E et al. (2019). The generation and use of recombinant extracellular vesicles as biological reference material. Nat Commun 10, 3288.
Geng T et al. (2021). Recent advancement and technical challenges in developing small extracellular vesicles for cancer drug delivery. Pharm Res 38, 179–197.
Gouin K et al. (2017). A comprehensive method for identification of suitable reference genes in extracellular vesicles. J Extracell Vesicles 6, 1347019.
Groot M and Lee H (2020). Sorting mechanisms for microRNAs into extracellular vesicles and their associated diseases. Cells 9, 1044.
Heath N et al. (2018). Rapid isolation and enrichment of extracellular vesicle preparations using anion exchange chromatography. Sci Rep 8, 5730.
Ilahibaks NF et al. (2024). Extracellular vesicles as vehicles for drug delivery to the heart. Eur Heart J, ehae099.
Irmer B et al. (2023). Extracellular vesicles in liquid biopsies as biomarkers for solid tumors. Cancers (Basel) 15, 1307.
Jia S et al. (2014). Emerging technologies in extracellular vesicle-based molecular diagnostics. Expert Rev Mol Diagn 14, 307–321.
Koch LF et al. (2024). Novel insights into the isolation of extracellular vesicles by anion exchange chromatography. Front Bioeng Biotechnol 11, 1298892.
Liangsupree T et al. (2021). Modern isolation and separation techniques for extracellular vesicles. J Chromatogr A 1636, 461773.
Lianidou E and Pantel K (2019). Liquid biopsies. Genes Chromosomes Cancer 58, 219–232.
Malvicini R et al. (2023). Ion exchange chromatography as a simple and scalable method to isolate biologically active small extracellular vesicles from conditioned media. PLoS One 18, e0291589.
Mateescu B et al. (2017). Obstacles and opportunities in the functional analysis of extracellular vesicle RNA — an ISEV position paper. J Extracell Vesicles 6, 1286095.
Maugeri M et al. (2019). Linkage between endosomal escape of LNP-mRNA and loading into EVs for transport to other cells. Nat Commun 10, 4333.
Newman LA et al. (2023). Extracellular Vesicle and contaminant markers in blood derivatives using multiple reaction monitoring. Methods Mol Biol 2628, 301–230.
Nieuwland R and Siljander PRM (2024). A beginner’s guide to study extracellular vesicles in human blood plasma and serum. J Extracell Vesicles 13, e12400.
Paolini L et al. (2016). Residual matrix from different separation techniques impacts exosome biological activity. Sci Rep 6, 23550.
Rayamajhi S et al. (2024). Extracellular vesicles as liquid biopsy biomarkers across the cancer journey: from early detection to recurrence. Clin Chem 70, 206–219.
Reiter K et al. (2019). Separation of virus-like particles and extracellular vesicles by flow-through and heparin affinity chromatography. J Chromatogr A 1588, 77–84.
Rodosthenous RS (2020). Profiling extracellular long RNA transcriptome in human plasma and extracellular vesicles for biomarker discovery. iScience 23, 101182.
Seo N et al. (2022). Distinguishing functional exosomes and other extracellular vesicles as a nucleic acid cargo by the anion‐exchange method. J Extracell Vesicles 11, e12205.
Sidhom K et al. (2020). A Review of exosomal isolation methods: is size exclusion chromatography the best option? Int J Mol Sci 21, 6466.
Simonsen JB (2017). What are we looking at? Extracellular vesicles, lipoproteins, or both? Circ Res 121, 920–922.
Sork H et al. (2018). Heterogeneity and interplay of the extracellular vesicle small RNA transcriptome and proteome. Sci Rep 8, 10813.
Théry C et al. (2018). Minimal information for studies of extracellular vesicles 2018 (MISEV2018): a position statement of the International Society for Extracellular Vesicles and update of the MISEV2014 guidelines. J Extracell Vesicles 7, 1535750.
Zeringer E et al. (2015). Strategies for isolation of exosomes. Cold Spring Harb Protoc 2015, 319–323.
Zhou Y et al. (2023). Large-scale heparin-based bind-and-elute chromatography identifies two biologically distinct populations of extracellular vesicles. J Extracell Vesicles 12, e12327.