R-loops are cellular three-stranded nucleic acid structures comprised of a DNA:RNA hybrid and a displaced DNA strand. The first R-loops, which can be formed co-transcriptionally in cis and post-transcriptionally in trans by Rad51 and Rad52, were visualized by electron microscopy in 1976 (Costantino and Koshland 2015, Thomas et al. 1976).
Opposite Sides of R-Loop Formation
In recent years, R-loops have been shown to be important in many biological processes; for example, they play a role in regulating gene expression (Ginno et al. 2013, Skourti-Stathaki et al. 2011, Skourti-Stathaki et al. 2014b), DNA methylation (Ginno et al. 2012), histone modifications (Castellano-Pozo et al. 2013), immunoglobulin class switch recombination (Yu et al. 2003), and driving embryonic stem cell differentiation (Chen et al. 2015). Interestingly, R-loops can also be formed at telomeres, which impacts telomere-length dynamics and senescence (Balk et al. 2013).
Based on their importance, it is no surprise that dysregulation of R-loop structures is associated with several human diseases, including neurodegenerative syndromes and cancer (Groh and Gromak 2014). R-loop formation has to be strictly regulated by the cell to ensure genome integrity, for example, the exposed ssDNA on R-loops is more vulnerable to breakage, deamination, and nuclease cleavage, resulting in DNA damage, mutations, and chromosome rearrangements (Constantino and Koshland 2015). In addition, R-loops can block replication fork progression, leading to double strand breaks (Gan et al. 2011). A number of DNA damage and repair proteins such as BRCA1 and members of the Fanconi Anemia protein family are known as important regulators of R-loop levels (Bhatia et al. 2014, Schwab et al. 2015).
How to Control Levels of R-Loop Formation?
As R-loop formation has to be tightly controlled, cells have mechanisms to prevent their generation. RNA processing factors and members of the topoisomerase family assist in the regulation of R-loop levels (Skourti-Stathaki et al. 2014a). Another mechanism for R-loop dissolution is activity of RNase H1 and RNase H2 enzymes, which specifically degrade the RNA in hybrids (Hiller et al. 2012). In addition, RNA-DNA helicases, such as probable helicase senataxin (SETX) and ATP-dependent RNA helicase A (DHX9), are reported to unwind the RNA/DNA hybrids thereby preventing R-loop accumulation (Skourti-Stathaki et al. 2011; Costantino and Koshland 2014; Cristini et al. 2018) (Figure 1). In addition, the 5′–3′ exonuclease XRN2 binds to and is associated with R-loop-mediated transcription termination (Skourti-Stathaki et al. 2011; Cristini et al. 2018).
The antibody-based method used for R-loop mapping is called DIP or DRIP (DNA/RNA immunoprecipitation) and it relies on the anti-DNA-RNA hybrid antibody, clone S9.6, used for the immunoprecipitation step (Boguslawski et al. 1986).
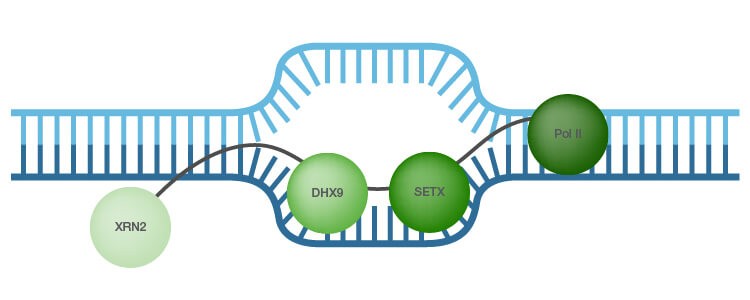
Fig. 1. R-loops. A recent R-loop interactome study identified ATP-dependent RNA helicase, also known as DHX9, as a top promoter of R-loop formation suppression (adapted from Cristini et al. 2018 and Skourti-Stathaki et al. 2011). The 5′–3′ exonuclease XRN2 is associated with R-loop-mediated transcription termination (Skourti-Stathaki et al. 2011). SETX, probable helicase senetaxin; Pol II, polymerase II.
R-Loops as Potential Therapeutic Targets
The potential of R-loops as therapeutic cancer targets is currently under investigation. This type of research has been fueled by studies showing the impact of the cancer drugs topotecan and camptothecin, inhibitors of topoisomerase, on R-loop formation in vivo (Powell et al. 2013, Marinello et al. 2013). In addition to its therapeutic potential, identification of small-molecule inhibitors of RNase H2 may serve as tools to investigate the function of R-loops (White et al. 2013). In fact, RNase H2 was identified in a genomic screen as a putative anti-cancer drug target (Flanagan et al. 2009). In addition to cancer, R-loops could also provide potential targets for treatment of neurological disorders, such as trinucleotide expansion diseases (Colak et al. 2014). There is also potential for the wider use of anti-DNA-RNA hybrid antibodies, which is supported by the use of the S9.6 clone for sensitive miRNA analysis (Qavi et al. 2011). Surprisingly, a small number of miRNAs (∼200 in total) can be sufficient for cancer classification, emphasizing the potential of miRNA profiling in cancer diagnosis (Lu et al. 2005).
Additional data support the conflicting nature of R loops — they can be detrimental structures, which when uncontrolled result in disease, but they can also have a positive effect by regulating essential cellular processes. The future challenge is to fully understand the complex mechanisms balancing these two opposing effects.
References
Balk B et al. (2013). Telomeric RNA-DNA hybrids affect telomere-length dynamics and senescence. Nat Struct Mol Biol 20, 1,199–1,205.
Bhatia V et al. (2014). BRCA2 prevents R-loop accumulation and associates with TREX-2 mRNA export factor PCID2. Nature 511, 362–365.
Boguslawski SJ et al. (1986). Characterization of monoclonal antibody to DNA. RNA and its application to immunodetection of hybrids. J Immunol Methods 1, 123–130.
Castellano-Pozo M et al. (2013). R loops are linked to histone H3 S10 phosphorylation and chromatin condensation. Mol Cell 52, 583–590.
Chen PB et al. (2015). R loops regulate promoter-proximal chromatin architecture and cellular differentiation. Nat Struct Mol Biol 22, 999–1,007.
Colak D et al. (2014). Promoter-bound trinucleotide repeat mRNA drives epigenetic silencing in Fragile X syndrome. Science 343, 1,002–1,005.
Costantino L and Koshland D (2015). The yin and yang of R-loop biology. Curr Opin Cell Biol 34, 39–45.
Cristini A et al. (2018). RNA/DNA hybrid interactome identifies DXH9 as a molecular player in transcriptional termination and R-Loop–associated DNA damage. Cell Rep 23, 1,891–1,905.
Flanagan JM et al. (2009). Genomics screen in transformed stem cells reveals RNASEH2A, PPAP2C, and ADARB1 as putative anticancer drug targets. Mol Cancer Ther 8, 249–260.
Gan W et al. (2011). R-loop-mediated genomic instability is caused by impairment of replication fork progression. Genes Dev 25, 2,041–2,056.
Ginno PA et al. (2012). R-loop formation is a distinctive characteristic of unmethylated human CpG island promoters. Mol Cell 45, 814–825.
Ginno PA et al. (2013). GC skew at the 5′ and 3′ ends of human genes links R-loop formation to epigenetic regulation and transcription termination. Genome Res 23, 1,590–1,600.
Groh M and Gromak N (2014). Out of balance: R-loops in human disease. PLoS Genet 10, e1004630.
Hiller B et al. (2012). Mammalian RNase H2 removes ribonucleotides from DNA to maintain genome integrity. J Exp Med 209, 1,419–1,426.
Lu J et al. (2005). MicroRNA expression profiles classify human cancers. Nature 9, 834–838.
Marinello J et al. (2013). Antisense transcripts enhanced by camptothecin at divergent CpG-island promoters associated with bursts of topoisomerase I–DNA cleavage complex and R-loop formation. Nucleic Acids Res 41, 10,110–10,123.
Powell WT et al. (2013). R-loop formation at Snord116 mediates topotecan inhibition of Ube3a-antisense and allele-specific chromatin decondensation. Proc Natl Acad Sci USA 110, 13,938–13,943.
Qavi AJ et al. (2011). Anti-DNA:RNA antibodies and silicon photonic microring resonators: increased sensitivity for multiplexed microRNA detection. Anal Chem 83, 5,949–5,956.
Schwab RA et al. (2015). The Fanconi anemia pathway maintains genome stability by coordinating replication and transcription. Mol Cell 60, 351–361.
Skourti-Stathaki K et al. (2011). Human senataxin resolves RNA/DNA hybrids formed at transcriptional pause sites to promote XRN2-dependent termination. Mol Cell 42, 794–805.
Skourti-Stathaki K et al. (2014a). A double-edged sword: R loops as threats to genome integrity and powerful regulators of gene expression. Genes Dev 28, 1,384–1,396.
Skourti-Stathaki K et al. (2014b). R-loops induce repressive chromatin marks over mammalian gene terminators. Nature 516, 436–439.
Thomas M et al. (1976). Hybridization of RNA to double-stranded DNA: formation of R-loops. Proc Natl Acad Sci USA 73, 2,294–2,298.
White R et al. (2013). Identification of small-molecule inhibitors of the ribonuclease H2 enzyme. J Biomol Screen 18, 610–620.
Yu K et al. (2003). R-loops at immunoglobulin class switch regions in the chromosomes of stimulated B cells. Nat Immunol 4, 442–451.
Bio-Rad is a trademark of Bio-Rad Laboratories, Inc. in certain jurisdictions.




