Abstract
In this study, triplex one-step reverse transcription quantitative PCR (RT-qPCR) assays have been developed for a panel of biomarkers relevant to colorectal cancer. The results using the triplex one-step RT-qPCR are equivalent to those obtained from slower and more costly singleplex assays. We show that implementing the multiplex one-step RT-qPCR can lead to significant savings in time and reagent costs relative to traditional singleplex RT-qPCR.
Introduction • Materials and Methods • Results and Discussion • Conclusions • References
Figures
Validation of Triplex RT-qPCR Assays
Fig.1. Validation of triplex assays for multiple targets.Comparison of Biomarker Expression between Cancerous and Normal Tissue Samples

Fig. 2. Differential biomarker expression in matched pairs of cancerous and adjacent normal tissue RNA samples.
Introduction
Colorectal cancer (CRC), the third most common cancer in both men and women, is the second leading cause of cancer mortality in the world. As a CRC neoplasm develops and progresses, its pattern of gene expression changes. Thus, differentially expressed genes can be used as biomarkers to assess and monitor CRC neoplasms (Chan et al. 2008, Jube et al. 2012, Nummela et al. 2012, Wu et al. 2012).
Clinical and translational research commonly relies on RT-qPCR (described in detail in the Applications & Technologies pages of the Bio-Rad website) for assessment of known or putative biomarkers to monitor disease. Such work is often limited by the large number of samples that need to be processed, the size of the biomarker panel, the high cost of the RT-qPCR reagents, and the time involved in setting up and running the experiments. One way to overcome these limitations is to make RT-qPCR faster and less expensive. Instead of performing reverse transcription and qPCR in two separate steps, the two reactions can be combined into a single one-step RT-qPCR reaction. Another approach is to analyze several target genes simultaneously in multiplex reactions. For further time and cost savings, one-step RT-qPCR and multiplexing can be combined into multiplex one-step RT-qPCR. In this case, an RNA sample is combined with a one-step RT-qPCR reagent and two or more probe-based target gene PCR assays. The mix then undergoes reverse transcription followed by multiplex qPCR in a real-time PCR instrument.
Here we demonstrate how to design and validate triplex one-step RT-qPCR assays using a panel of biomarkers that is important to CRC research. We show that triplex one-step RT-qPCR results obtained with Bio-Rad’s iTaq™ Universal Probes One-Step Kit are equivalent to results obtained from slower and more costly singleplex analysis. This suggests that multiplex one-step RT-qPCR assays can be employed as a time- and cost-saving alternative to traditional qPCR protocols.
Materials and Methods
Samples
We obtained total RNA, corresponding to matched colorectal tumor tissue and adjacent normal tissue, from two patients through a commercial source (BioChain Institute, Inc.). The samples were from (1) a 68-year-old male with poorly differentiated adenocarcinoma and (2) a 62-year-old female with moderately differentiated adenocarcinoma.
Target Genes
Four target genes that exhibit differential expression in CRC samples were assessed (TGFBI, COL1A1, HMGB1, and COL3A1). Amplification primers and probes were designed using Primer3Plus (www.bioinformatics.nl/cgi-bin/primer3plus/primer3plus.cgi/) and synthesized by Integrated DNA Technologies, Inc. (IDT). GAPDH primers and probe were from Biosearch Technologies, Inc. Table 1 lists the primer and probe sequences.
Table 1. Primer and probe information.

Triplex Reactions
We developed two sets of triplex assays. Set 1 contains TGFBI, COL1A1, and GAPDH. Set 2 contains HMGB1, COL3A1, and GAPDH. GAPDH serves as an internal reference gene control in both triplex assays.
One-Step RT-qPCR
One-step RT-qPCR was performed using the iTaq Universal Probes One-Step Kit as directed by the manufacturer on a CFX96™ Real-Time PCR Detection System (Bio-Rad). For all target genes, primers and probes were used at 250 nM final concentrations. For the GAPDH reference gene, primers were used at 100 nM and the probe at 250 nM final concentrations. The GAPDH assay included a lesser amount of primers because GAPDH is highly expressed; lowering the concentration of GAPDH primers prevents GAPDH amplification from depleting reaction components. Cycling conditions were 10 min at 50°C (reverse transcription), 3 min at 95°C followed by 40 cycles of 10 sec at 95°C, and 1 min at 60°C.
Triplex Assay Validation
To validate the triplex assays, triplex and singleplex one-step RT-qPCR performance was compared using tenfold serial dilutions of HeLa RNA (100 ng–10 pg). The experiment was repeated three times independently.
Analysis of CRC Samples
Each patient RNA sample (1 ng) was analyzed by one-step RT-qPCR in both triplex and singleplex. Each sample was analyzed four times independently.
Results and Discussion
Triplex Assay Validation
We chose to analyze a panel of four biomarkers that are reported to be overexpressed in CRC. COL1A1, COL3A1, and TGFBI act in the extracellular matrix receptor interaction pathway (Chan et al. 2008, Nummela et al. 2012, Wu et al. 2012), whereas HMGB1 has potential roles in tumor cell survival and metastasis (Jube et al. 2012).
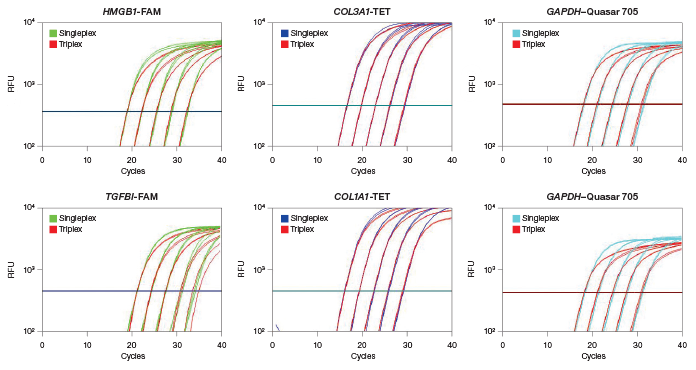
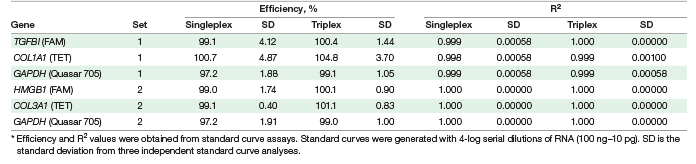
We developed two sets of triplex assays. Each set contains two CRC biomarkers and the GAPDH reference gene that serves as an internal normalization control (Table 1). To validate the triplex assays in one-step RT-qPCR, we compared triplex performance with singleplex performance using serially diluted HeLa RNA. We found that the triplex assays and singleplex assays give equivalent performance. At all input RNA levels, the quantification cycle (Cq) values of the triplex and singleplex reactions were virtually indistinguishable (Figure 1) and the qPCR efficiency and R2 values were statistically the same (Table 2). These data indicate that triplex assays can be performed instead of singleplex assays without a loss in data quality.
Fig. 1. Validation of triplex assays for multiple targets. Amplification plots of tenfold serial RNA dilutions are shown (100 ng–10 pg). For all targets, the Cq values for the singleplex and triplex reactions are virtually indistinguishable, indicating that triplex reactions perform as well as singleplex reactions. RFU, relative fluorescence units.
Table 2. Comparison of singleplex and triplex one-step RT-qPCR reactions.*
Analysis of Samples from Patients with CRC
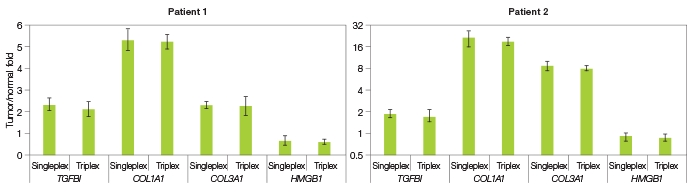
To confirm that triplex and singleplex assays yield equivalent results we analyzed RNA samples from two patients with CRC. We obtained two RNA samples from each patient: one sample from a malignant colon region and the other sample from an adjacent colon region that was not cancerous. We then quantified the expression of each biomarker using triplex and singleplex one-step RT-qPCR, normalized biomarker expression with GAPDH, and determined whether each biomarker exhibits differential expression in cancerous tissue. Our results show that, in the two patients analyzed, TGFB1, COL1A1, and COL3A1 are upregulated in cancerous tissue while HMGB1 is not (Figure 2). Importantly, the triplex and singleplex assays produce results that are statistically equivalent. These findings confirm that our triplex assays can substitute for singleplex assays without compromising the quality of results. This implies that in future work, patient samples can be reliably analyzed by triplex one-step RT-qPCR.
Fig. 2. Differential biomarker expression in matched pairs of cancerous and adjacent normal tissue RNA samples. Gene expression levels in RNA samples from two patients with CRC were determined using triplex and singleplex one-step RT-qPCR. Biomarker expression was normalized using the GAPDH internal reference gene control. The ratio of biomarker expression in cancerous tissue relative to adjacent normal tissue is shown for each patient. The experiment was repeated four times; error bars represent the 95% confidence interval.
Benefits of Multiplex One-Step RT-qPCR
Our work demonstrates that properly validated one-step RT-qPCR triplex reactions using Bio-Rad’s iTaq Universal Probes One-Step Kit generate results that are equivalent to singleplex reactions. Implementing multiplex one-step RT-qPCR can lead to significant savings in time and reagent costs relative to traditional singleplex RT-qPCR. Multiplex one-step RT-qPCR can also lead to improved data quality since researchers need to perform only one reaction using a single reagent. This saves time and can lead to better results because there is no chance of manipulation error between the reverse transcription and qPCR steps. Multiplex analysis also reduces sample utilization because fewer reactions need to be processed. Lastly, because each multiplex reaction can be internally controlled, with GAPDH in our example, there will be less variability in experiments involving relative gene expression.
Conclusions
Multiplex one-step RT-qPCR provides an excellent alternative to traditional RT-qPCR protocols, especially when time, sample, and cost are limited.
References
Chan SK et al. (2008). Meta-analysis of colorectal cancer gene expression profiling studies identifies consistently reported candidate biomarkers. Cancer Epidemiol Biomarkers Prev 17, 543–552.
Jube S et al. (2012). Cancer cell secretion of the DAMP protein HMGB1 supports progression in malignant mesothelioma. Cancer Res 72, 3290–3301.
Nummela P et al. (2012). Transforming growth factor beta-induced (TGFBI) is an anti-adhesive protein regulating the invasive growth of melanoma cells. Am J Pathol 180, 1663–1674.
Wu Y et al. (2012). Transcriptome profiling of the cancer, adjacent non-tumor and distant normal tissues from a colorectal cancer patient by deep sequencing. PLoS One 7, e41001.
Visit bio-rad.com/web/1stepRTqPCR for more information.
FAM and TET are trademarks of Applera Corporation. Quasar is a trademark of Biosearch Technologies, Inc.
Myriad RBM is a trademark of Myriad RBM, Inc.
Purchase of iTaq Universal Probes One-Step Kit includes an immunity from suit under patents specified in the product insert to use only the amount purchased for the purchaser’s own internal research. No other patent rights are conveyed expressly, by implication, or by estoppel. Further information on purchasing licenses may be obtained by contacting the Director of Licensing, Applied Biosystems, 850 Lincoln Centre Drive, Foster City, California 94404, USA.