Any transcript. Any sample. Any amount.
The SEQuoia Complete Stranded RNA Library Prep Kit is the newest addition to Bio-Rad’s next-generation sequencing (NGS) space. The high-performance kit captures both long and short RNAs in a single library, even from limited and low-quality samples. Featuring SEQzyme, a proprietary engineered enzyme that couples cDNA synthesis with adaptor addition in a one-tube continuous synthesis reaction, this kit allows construction of libraries that are >99% stranded and suitable for next-generation sequencing on Illumina® platforms.
See why the SEQuoia Complete Kit is your top choice for high-performance RNA-Seq library prep.
Validated with FFPE and other low-quality samples. Gene discovery from limited formalin-fixed paraffin-embedded (FFPE) samples exhibits high concordance with matched frozen samples. 85% of genes identified in SEQuoia libraries from FFPE samples were also found in libraries from frozen samples, highlighting the biological relevance of the FFPE sequencing results obtained using the SEQuoia Complete Stranded RNA Library Prep Kit.
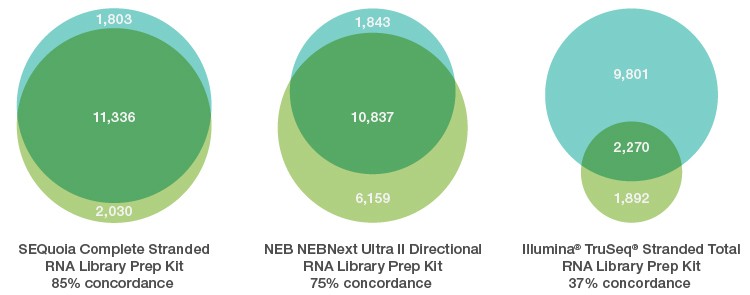
Gene discovery from limited FFPE samples exhibits high concordance with matched frozen samples using the SEQuoia Complete RNA Stranded Library Prep Kit. Overlap (■) in genes detected at a read depth of 1 RPKM in libraries constructed from 10 ng of rRNA-depleted Frozen (■) or FFPE (■) Matched Pair Total RNA: Human Adult Normal Tissue: Liver (BioChain catalog #R8234149-FP) using the SEQuoia Complete Stranded RNA Library Prep Kit or other commercial kits. Eighty-five percent of genes identified in SEQuoia libraries from the FFPE sample were also found in libraries from the frozen sample, highlighting the biological relevance of the FFPE sequencing results obtained using the SEQuoia Complete Stranded RNA Library Prep Kit.
Captures long and short RNAs in a single prep. The unique enzymatic properties of SEQzyme effectively capture fragments from highly degraded RNA and short RNA fragments (for example, snoRNA, miRNA, and tRNA), which are commonly missed by other commercial RNA library prep kits. The efficient simultaneous capture of diverse RNA fragments results in a library that more accurately represents the complete transcriptome.
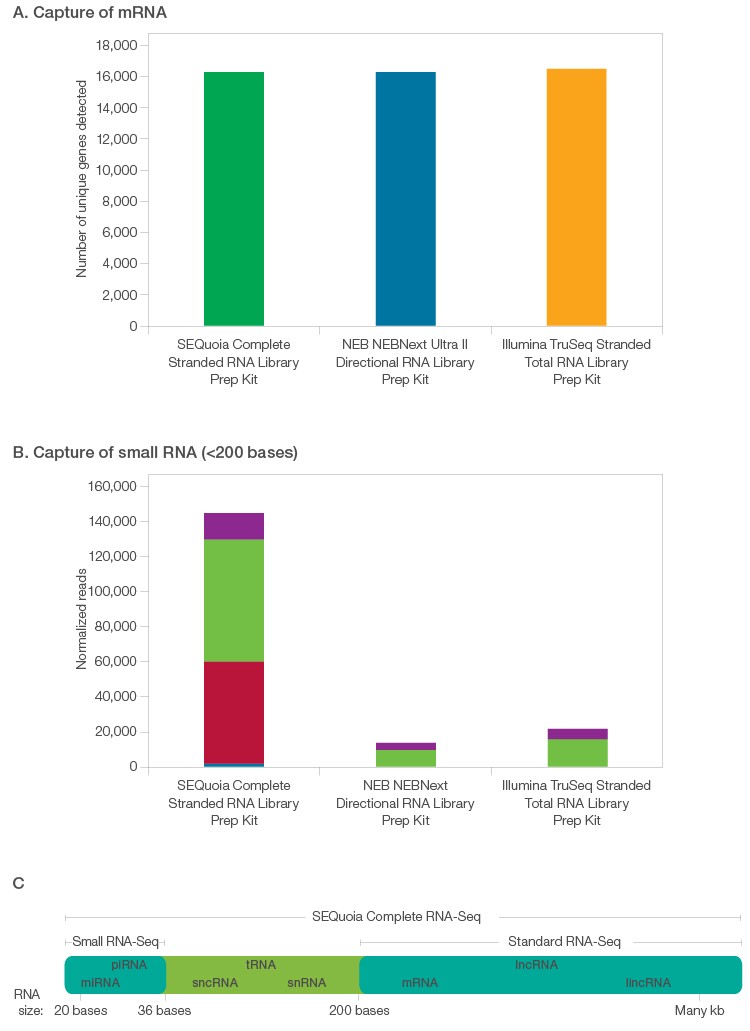
Libraries prepared using the SEQuoia Complete Stranded RNA Library Prep Kit produce richer datasets. Libraries were prepared with either the SEQuoia Complete Stranded RNA Library Prep Kit or other commercial kits using 10 ng of rRNA-depleted RNA and sequenced to a read depth of 10 million reads. A, the number of genes detected at >5 RPKM in libraries prepared using the SEQuoia Complete Stranded RNA Library Prep Kit was equivalent to that of libraries prepared using other kits. B, in addition to effectively capturing mRNA transcripts, libraries prepared with the SEQuoia Complete Stranded RNA Library Prep Kit included a far greater number of unique reads mapping to RNAs shorter than 200 nucleotides in length, including miRNA (■), tRNA (■), snoRNA (■), and snRNA (■), indicating a richer, more complex library than other commercial kits. C, the SEQuoia Complete Stranded RNA Library Prep Kit captures a greater diversity of RNA species than standard or small RNA-Seq kits combined.
Performs well with low sample input. The comprehensive, diverse libraries constructed from a wide range of starting materials produces substantially higher yields with fewer PCR cycles than other commercial kits when low quantities of degraded RNA are used as input.
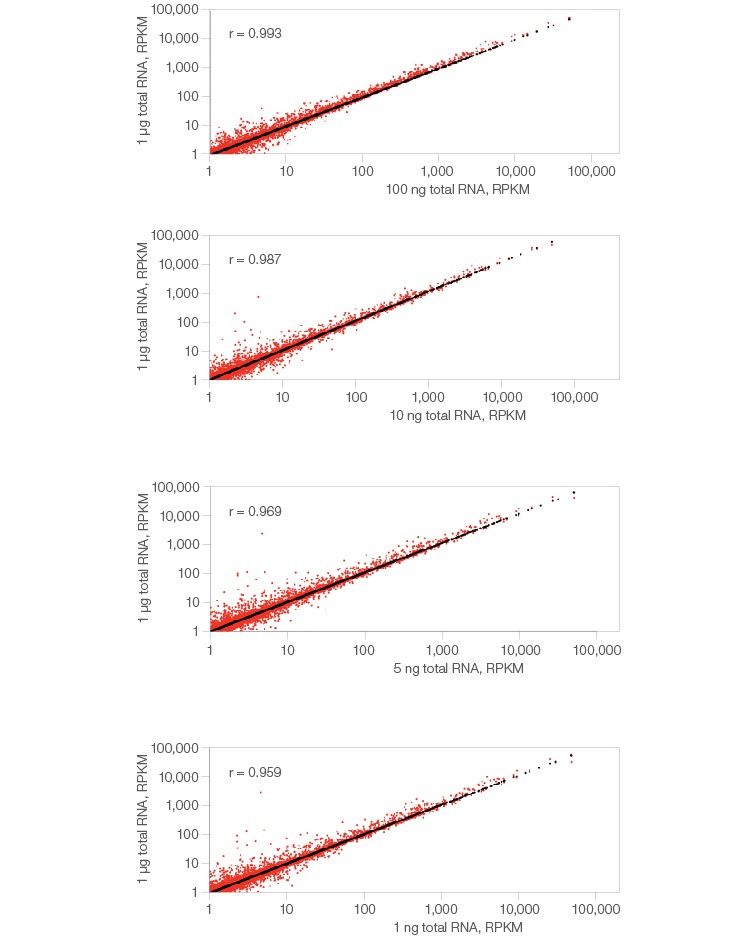
The SEQuoia Complete Stranded RNA Library Prep Kit produces libraries with consistent gene detection across a broad range of input RNA quantities. Varying amounts of Human Placenta Total RNA (Thermo Fisher Scientific Inc. #AM7950) ranging from 1 μg to 1 ng were rRNA depleted (NEBNext rRNA Depletion Kit, NEB catalog #E6310), used to construct libraries, and then sequenced. The numbers of reads per kilobase of transcript per million mapped reads (RPKM) derived from each library are plotted against RPKM derived from the 1 μg library (y-axis). Correlation between the libraries, calculated as the Pearson correlation coefficient (r), indicates exceptional detection concordance between the different input amounts, even with limited input.
Enables an efficient and streamlined library prep workflow. The kit is optimized to facilitate efficient library synthesis using a wide range of input RNA (100 pg to 1 μg) and is compatible with automated next-generation sequencing library construction. The streamlined workflow minimizes the number of pipetting steps and greatly reduces the overall protocol time to less than 4 hours.
Experience the power of the SEQuoia Complete Kit today.